You cannot select more than 25 topics
Topics must start with a letter or number, can include dashes ('-') and can be up to 35 characters long.
|
|
2 months ago | |
|---|---|---|
| main_files | 2 months ago | |
| Readme.md | 2 months ago | |
| iris_basic.csv | 2 months ago | |
| main.ipynb | 2 months ago | |
Readme.md
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv("iris_basic.csv")
print(df.head())
sl sw pl pw target tNames
0 5.1 3.5 1.4 0.2 0 setosa
1 4.9 3.0 1.4 0.2 0 setosa
2 4.7 3.2 1.3 0.2 0 setosa
3 4.6 3.1 1.5 0.2 0 setosa
4 5.0 3.6 1.4 0.2 0 setosa
x = df["pw"].to_numpy().reshape(-1, 1) # (150,1)
x
array([[0.2],
[0.2],
[0.2],
[0.2],
[0.2],
[0.4],
[0.3],
[0.2],
[0.2],
[0.1],
[0.2],
[0.2],
[0.1],
[0.1],
[0.2],
[0.4],
[0.4],
[0.3],
[0.3],
[0.3],
[0.2],
[0.4],
[0.2],
[0.5],
[0.2],
[0.2],
[0.4],
[0.2],
[0.2],
[0.2],
[0.2],
[0.4],
[0.1],
[0.2],
[0.2],
[0.2],
[0.2],
[0.1],
[0.2],
[0.2],
[0.3],
[0.3],
[0.2],
[0.6],
[0.4],
[0.3],
[0.2],
[0.2],
[0.2],
[0.2],
[1.4],
[1.5],
[1.5],
[1.3],
[1.5],
[1.3],
[1.6],
[1. ],
[1.3],
[1.4],
[1. ],
[1.5],
[1. ],
[1.4],
[1.3],
[1.4],
[1.5],
[1. ],
[1.5],
[1.1],
[1.8],
[1.3],
[1.5],
[1.2],
[1.3],
[1.4],
[1.4],
[1.7],
[1.5],
[1. ],
[1.1],
[1. ],
[1.2],
[1.6],
[1.5],
[1.6],
[1.5],
[1.3],
[1.3],
[1.3],
[1.2],
[1.4],
[1.2],
[1. ],
[1.3],
[1.2],
[1.3],
[1.3],
[1.1],
[1.3],
[2.5],
[1.9],
[2.1],
[1.8],
[2.2],
[2.1],
[1.7],
[1.8],
[1.8],
[2.5],
[2. ],
[1.9],
[2.1],
[2. ],
[2.4],
[2.3],
[1.8],
[2.2],
[2.3],
[1.5],
[2.3],
[2. ],
[2. ],
[1.8],
[2.1],
[1.8],
[1.8],
[1.8],
[2.1],
[1.6],
[1.9],
[2. ],
[2.2],
[1.5],
[1.4],
[2.3],
[2.4],
[1.8],
[1.8],
[2.1],
[2.4],
[2.3],
[1.9],
[2.3],
[2.5],
[2.3],
[1.9],
[2. ],
[2.3],
[1.8]])
y = df["target"].to_numpy().reshape(-1, 1) # (150,1)
y = (y == 0).astype(float)
y
array([[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[1.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.],
[0.]])
def sigmoid(z):
z = np.clip(z, -500, 500)
sig = 1.0 / (1.0 + np.exp(-z))
return sig
def log_loss(y, p, eps=1e-12):
p = np.clip(p, eps, 1 - eps)
return -np.mean(y*np.log(p) + (1-y)*np.log(1-p))
lr=0.1
epochs=2000
l2=0.0,
X = np.column_stack([x, np.ones_like(x)])
m = X.shape[0]
theta = np.zeros((2,1))
theta
array([[0.],
[0.]])
X.T
array([[0.2, 0.2, 0.2, 0.2, 0.2, 0.4, 0.3, 0.2, 0.2, 0.1, 0.2, 0.2, 0.1,
0.1, 0.2, 0.4, 0.4, 0.3, 0.3, 0.3, 0.2, 0.4, 0.2, 0.5, 0.2, 0.2,
0.4, 0.2, 0.2, 0.2, 0.2, 0.4, 0.1, 0.2, 0.2, 0.2, 0.2, 0.1, 0.2,
0.2, 0.3, 0.3, 0.2, 0.6, 0.4, 0.3, 0.2, 0.2, 0.2, 0.2, 1.4, 1.5,
1.5, 1.3, 1.5, 1.3, 1.6, 1. , 1.3, 1.4, 1. , 1.5, 1. , 1.4, 1.3,
1.4, 1.5, 1. , 1.5, 1.1, 1.8, 1.3, 1.5, 1.2, 1.3, 1.4, 1.4, 1.7,
1.5, 1. , 1.1, 1. , 1.2, 1.6, 1.5, 1.6, 1.5, 1.3, 1.3, 1.3, 1.2,
1.4, 1.2, 1. , 1.3, 1.2, 1.3, 1.3, 1.1, 1.3, 2.5, 1.9, 2.1, 1.8,
2.2, 2.1, 1.7, 1.8, 1.8, 2.5, 2. , 1.9, 2.1, 2. , 2.4, 2.3, 1.8,
2.2, 2.3, 1.5, 2.3, 2. , 2. , 1.8, 2.1, 1.8, 1.8, 1.8, 2.1, 1.6,
1.9, 2. , 2.2, 1.5, 1.4, 2.3, 2.4, 1.8, 1.8, 2.1, 2.4, 2.3, 1.9,
2.3, 2.5, 2.3, 1.9, 2. , 2.3, 1.8],
[1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. , 1. ,
1. , 1. , 1. , 1. , 1. , 1. , 1. ]])
for i in range(epochs):
z = X @ theta # (m,1)
h = sigmoid(z) # (m,1)
grad = (X.T @ (h - y)) / m # (2,1) <-- from your formula
theta -= lr * grad
#if (i % 0 == 0 or t == epochs-1):
# print(f"{i:4d} loss={log_loss(y, h):.6f} w={theta[0,0]:.6f} b={theta[1,0]:.6f}")
w, b = theta[0,0], theta[1,0]
def predict_proba(x, w, b):
x = np.asarray(x, float).reshape(-1)
return sigmoid(w*x + b)
def predict(x, w, b, thresh=0.5):
return (predict_proba(x, w, b) >= thresh).astype(int)
rng = np.random.default_rng(0)
m = 120
xNew = np.linspace(-0.5, 2.5, m)
p = predict_proba(xNew, w, b)
print(f"\nLearned: w={w:.3f}, b={b:.3f}, loss={log_loss(p.reshape(-1,1), p.reshape(-1,1)):.4f}")
Learned: w=-5.989, b=4.279, loss=0.1812
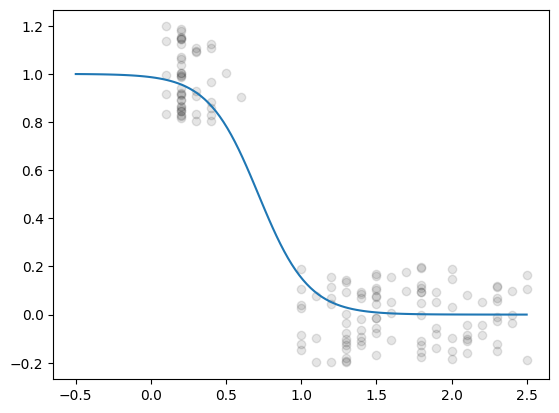
yJitter = y +np.random.uniform(-0.2, 0.2, size=y.shape)
plt.plot(x, yJitter, 'ok', alpha=0.1)
plt.plot(xNew,p)
[<matplotlib.lines.Line2D at 0x112e5cd70>]
Multi-Parametric Binary Classifier
x1 = df["sl"].to_numpy()
x2 = df["sw"].to_numpy()
X = np.column_stack([ np.ones_like(x1), x1, x2])
y = df["target"].to_numpy()
#y = (y == 2).astype(float)
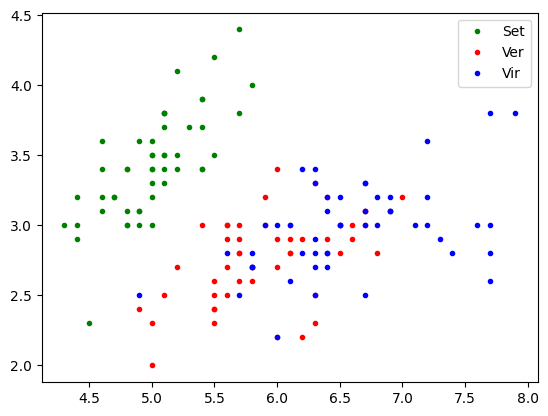
plt.plot(x1[y==0], x2[y==0],'.g' ,label='Set')
plt.plot(x1[y==1], x2[y==1],'.r', label='Ver')
plt.plot(x1[y==2], x2[y==2],'.b', label='Vir')
plt.legend()
plt.show()
y = df["target"].to_numpy()
y = (y == 2).astype(float)
def predict_proba(X, theta):
z = X@theta
return sigmoid(z)
lr=0.01
epochs=5000
m = X.shape[0]
theta = np.random.randn(3)
theta
array([ 0.68799477, 0.14542591, -0.00438461])
for i in range(epochs):
z = X @ theta # (m,1)
h = 1/(1+np.exp(-z)) # (m,1)
grad = (X.T @ (h - y)) / m # (2,1) <-- from your formula
theta -= lr * grad
if (i % 100 == 0):
print(f"{i:4d} loss={log_loss(y, h):.6f}")
theta
0 loss=1.178088
100 loss=0.647786
200 loss=0.630318
300 loss=0.615215
400 loss=0.602111
500 loss=0.590698
600 loss=0.580716
700 loss=0.571949
800 loss=0.564217
900 loss=0.557369
1000 loss=0.551279
1100 loss=0.545841
1200 loss=0.540968
1300 loss=0.536583
1400 loss=0.532624
1500 loss=0.529037
1600 loss=0.525776
1700 loss=0.522803
1800 loss=0.520083
1900 loss=0.517587
2000 loss=0.515291
2100 loss=0.513174
2200 loss=0.511214
2300 loss=0.509398
2400 loss=0.507709
2500 loss=0.506136
2600 loss=0.504667
2700 loss=0.503292
2800 loss=0.502002
2900 loss=0.500790
3000 loss=0.499649
3100 loss=0.498572
3200 loss=0.497554
3300 loss=0.496590
3400 loss=0.495676
3500 loss=0.494807
3600 loss=0.493980
3700 loss=0.493191
3800 loss=0.492438
3900 loss=0.491718
4000 loss=0.491028
4100 loss=0.490367
4200 loss=0.489731
4300 loss=0.489120
4400 loss=0.488532
4500 loss=0.487964
4600 loss=0.487416
4700 loss=0.486887
4800 loss=0.486374
4900 loss=0.485877
array([-0.45875015, 1.02970203, -2.10588172])
x1New, x2New = np.meshgrid(
np.linspace(3,8,100).reshape(-1,1),
np.linspace(0,6,100).reshape(-1,1))
XNew = np.column_stack([np.ones(x1New.size), x1New.ravel(), x2New.ravel()])
z = XNew @ theta
yPred = 1 / (1 + np.exp(-z))
zz = yPred.reshape(x1New.shape)
zz
array([[9.32789869e-01, 9.35977755e-01, 9.39024319e-01, ...,
9.99535858e-01, 9.99559369e-01, 9.99581689e-01],
[9.24332755e-01, 9.27890766e-01, 9.31293909e-01, ...,
9.99472707e-01, 9.99499415e-01, 9.99524770e-01],
[9.14908551e-01, 9.18870810e-01, 9.22664164e-01, ...,
9.99400969e-01, 9.99431308e-01, 9.99460111e-01],
...,
[5.83101559e-05, 6.14226291e-05, 6.47012289e-05, ...,
8.96718270e-03, 9.44134205e-03, 9.94032212e-03],
[5.13237738e-05, 5.40633488e-05, 5.69491493e-05, ...,
7.90122160e-03, 8.31948909e-03, 8.75970294e-03],
[4.51744213e-05, 4.75857700e-05, 5.01258268e-05, ...,
6.96108505e-03, 7.32995218e-03, 7.71821361e-03]], shape=(100, 100))
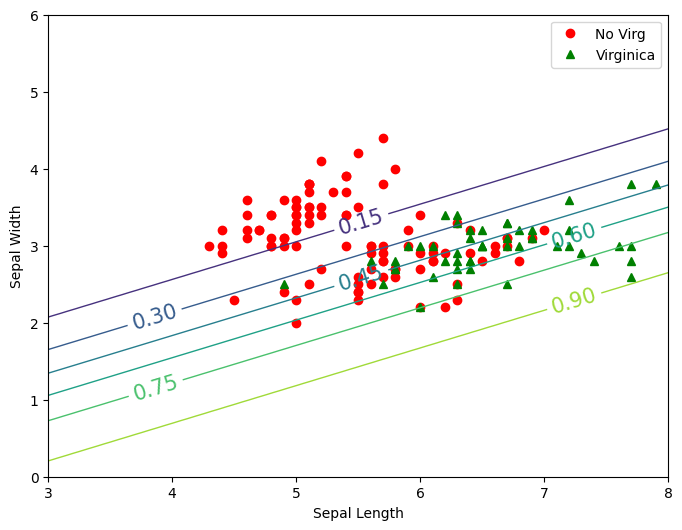
plt.figure(figsize=(8,6))
plt.plot(x1[y==0], x2[y==0],'or' ,label='No Virg')
plt.plot(x1[y==1], x2[y==1],'g^',label='Virginica')
contour = plt.contour(x1New,x2New,zz, linewidths=1)
plt.clabel(contour, inline=1,fontsize=15)
plt.xlabel("Sepal Length")
plt.ylabel("Sepal Width")
plt.legend()
plt.show()
# Softmax model ???